def generate_synthetic_data(n_samples=500, ts_length=80, img_size=(32, 32)):
"""
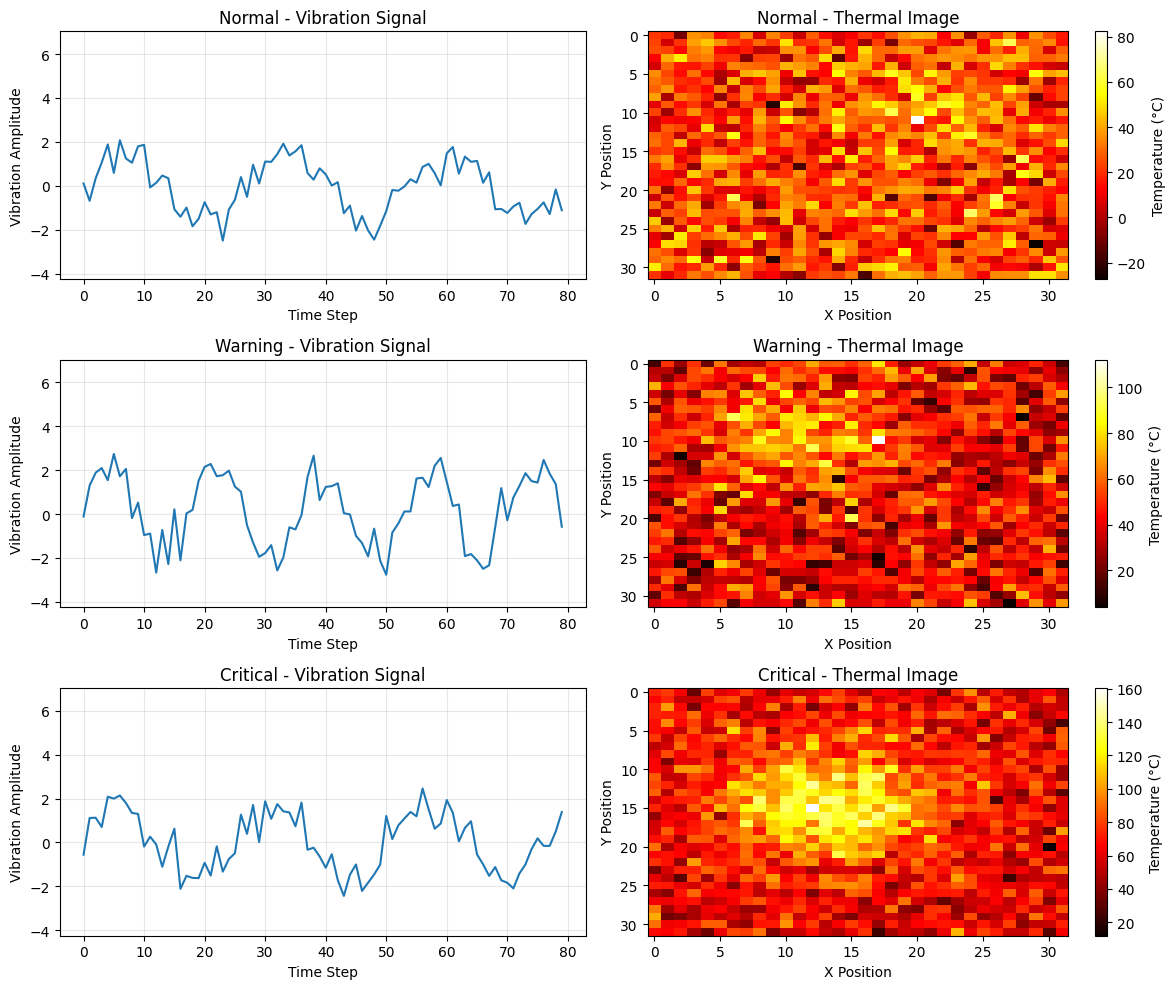
Generate synthetic multimodal data for equipment monitoring.
This function creates ambiguous data where each modality alone is insufficient,
but combining both provides better classification. Two types of issues are simulated:
- Type A: High vibration (looks critical) but low temperature (actually warning)
- Type B: Low vibration (looks normal) but high temperature (actually warning)
Returns:
time_series: vibration sensor data (n_samples, ts_length)
images: thermal camera data (n_samples, img_size[0], img_size[1])
labels: equipment health status (n_samples,)
"""
time_series = []
images = []
labels = []
for _ in range(n_samples):
# Randomly assign a health status Normal, Warning, Critical
label = np.random.choice([0, 1, 2], p=[0.45, 0.40, 0.15])
labels.append(label)
ambiguous_type = None
if label == 1 and np.random.random() < 0.85:
ambiguous_type = np.random.choice(["A", "B"])
elif label == 0 and np.random.random() < 0.40:
ambiguous_type = "slight_elevation" # Many normals look elevated
elif label == 2 and np.random.random() < 0.35:
ambiguous_type = "one_misleading" # Many criticals look normal in one modality
# Generate time series with high overlap - amplitude and frequency ranges heavily overlap
if label == 0: # Normal
if ambiguous_type == "slight_elevation":
# Overlaps strongly with warning
t = np.linspace(0, np.random.uniform(5, 8) * np.pi, ts_length)
ts = np.random.uniform(1.2, 1.6) * np.sin(t) + np.random.normal(0, 0.5, ts_length)
base_temp = 25 + np.random.normal(0, 7)
else:
t = np.linspace(0, np.random.uniform(4, 7) * np.pi, ts_length)
ts = np.random.uniform(0.8, 1.3) * np.sin(t) + np.random.normal(0, 0.5, ts_length)
base_temp = 22 + np.random.normal(0, 8)
elif label == 1: # Warning - extremely ambiguous in time series
if ambiguous_type == "A":
# Type A: Looks critical in vibration but moderate temperature
t = np.linspace(0, np.random.uniform(9, 12) * np.pi, ts_length)
ts = np.random.uniform(1.8, 2.3) * np.sin(t) + np.random.normal(0, 0.6, ts_length)
# Add spikes that make it look critical
spike_indices = np.random.choice(ts_length, size=np.random.randint(3, 6), replace=False)
ts[spike_indices] += np.random.uniform(1, 2.5, size=len(spike_indices))
base_temp = 40 + np.random.normal(0, 9) # Moderate temp distinguishes it
elif ambiguous_type == "B":
# Type B: Looks normal in vibration but elevated temperature
t = np.linspace(0, np.random.uniform(4, 7) * np.pi, ts_length)
ts = np.random.uniform(1.0, 1.4) * np.sin(t) + np.random.normal(0, 0.5, ts_length)
base_temp = 52 + np.random.normal(0, 9) # High temp distinguishes it
else:
# Regular warning - still overlaps with both normal and critical
t = np.linspace(0, np.random.uniform(6, 9) * np.pi, ts_length)
ts = np.random.uniform(1.3, 1.8) * np.sin(t) + np.random.normal(0, 0.6, ts_length)
base_temp = 43 + np.random.normal(0, 10)
else: # Critical
if ambiguous_type == "one_misleading":
# One modality looks much less critical
if np.random.random() < 0.5:
# Lower vibration (overlaps with warning/normal) but very high temp
t = np.linspace(0, np.random.uniform(6, 9) * np.pi, ts_length)
ts = np.random.uniform(1.4, 1.9) * np.sin(t) + np.random.normal(0, 0.6, ts_length)
spike_indices = np.random.choice(ts_length, size=np.random.randint(1, 3), replace=False)
ts[spike_indices] += np.random.uniform(0.5, 1.5, size=len(spike_indices))
base_temp = 70 + np.random.normal(0, 8) # Temperature is clearly critical
else:
# Higher vibration but moderate temp (overlaps with warning)
t = np.linspace(0, np.random.uniform(10, 13) * np.pi, ts_length)
ts = np.random.uniform(1.9, 2.3) * np.sin(t) + np.random.normal(0, 0.7, ts_length)
spike_indices = np.random.choice(ts_length, size=np.random.randint(4, 7), replace=False)
ts[spike_indices] += np.random.uniform(1.5, 3, size=len(spike_indices))
base_temp = 58 + np.random.normal(0, 8) # Moderate temp
else:
# More typical critical but with variability
t = np.linspace(0, np.random.uniform(10, 14) * np.pi, ts_length)
ts = np.random.uniform(1.8, 2.4) * np.sin(t) + np.random.normal(0, 0.7, ts_length)
spike_indices = np.random.choice(ts_length, size=np.random.randint(4, 8), replace=False)
ts[spike_indices] += np.random.uniform(1.5, 3.5, size=len(spike_indices))
base_temp = 68 + np.random.normal(0, 10)
time_series.append(ts)
# Generate thermal image with HIGH ambiguity to make classification harder
# Much more overlap between classes to reduce image-only accuracy
img = np.random.normal(base_temp, 15, img_size) # Even higher noise
# Add hot spots with heavily overlapping patterns to create strong ambiguity
if label == 0: # Normal - but frequently looks like warning or critical
if np.random.random() < 0.55:
# Often looks like warning or critical
n_hotspots = np.random.randint(2, 4)
hotspot_intensity_factor = np.random.uniform(1.0, 1.8)
hotspot_base = 18
else:
n_hotspots = np.random.randint(1, 3)
hotspot_intensity_factor = np.random.uniform(0.7, 1.3)
hotspot_base = 12
elif label == 1: # Warning - highly ambiguous, overlaps heavily with both
if np.random.random() < 0.45:
# Looks like normal
n_hotspots = np.random.randint(1, 3)
hotspot_intensity_factor = np.random.uniform(0.7, 1.3)
hotspot_base = 14
elif np.random.random() < 0.45:
# Looks like critical
n_hotspots = np.random.randint(2, 4)
hotspot_intensity_factor = np.random.uniform(1.5, 2.2)
hotspot_base = 28
else:
n_hotspots = np.random.randint(2, 3)
hotspot_intensity_factor = np.random.uniform(1.0, 1.6)
hotspot_base = 20
else: # Critical - but often looks like warning or normal
if np.random.random() < 0.50:
# Frequently looks like warning or normal
n_hotspots = np.random.randint(2, 3)
hotspot_intensity_factor = np.random.uniform(1.1, 1.7)
hotspot_base = 22
else:
n_hotspots = np.random.randint(2, 4)
hotspot_intensity_factor = np.random.uniform(1.6, 2.4)
hotspot_base = 30
for _ in range(n_hotspots):
x, y = np.random.randint(6, img_size[0] - 6), np.random.randint(6, img_size[1] - 6)
intensity = hotspot_base + np.random.uniform(5, 25) * hotspot_intensity_factor # More variable intensity
# Create a Gaussian hot spot with more variable size
sigma = np.random.uniform(2.0, 7.0) # Wider range for hotspot size
xx, yy = np.meshgrid(np.arange(img_size[0]), np.arange(img_size[1]))
hotspot = intensity * np.exp(-((xx - x) ** 2 + (yy - y) ** 2) / (2 * sigma**2))
img += hotspot.T
images.append(img)
return np.array(time_series), np.array(images), np.array(labels)
# Generate data
ts_data, img_data, labels = generate_synthetic_data()
print(f"Time series shape: {ts_data.shape}")
print(f"Images shape: {img_data.shape}")
print(f"Labels shape: {labels.shape}")
print()
print("Class distribution:")
print(f" Normal: {np.sum(labels == 0)} ({np.sum(labels == 0) / len(labels) * 100:.1f}%)")
print(f" Warning: {np.sum(labels == 1)} ({np.sum(labels == 1) / len(labels) * 100:.1f}%)")
print(f" Critical: {np.sum(labels == 2)} ({np.sum(labels == 2) / len(labels) * 100:.1f}%)")